
Now the same alignment with Shading and Trimmed turned off ĭon’t forget the Dots button too. Note the greyed out residues in the outlined areas Here is an alignment with Shading and Trimmed both turned ON. However, you can choose to completely hide those residues by turning off the Trimmed button. The Trimmed button controls the visibility of trimmed (or “clipped”) residues – these are residues that do not align to the reference sequence and so are greyed out to indicate that, and they are not included in consensus calculations. Coloring can be toggled on and off using the Shading button. The scale ranges from a dark red for poor quality, through white for “OK” quality (phred score 20 for an individual read) through dark green for high quality (phred score 40 or above). The Shading button turns on background coloring of the residues in the upper pane, based on quality values (these can be from Sanger reads or from NGS reads). This.MacVector’s Align to Reference Editor and the Contig Editor in Assembly Projects have two useful functions for visualizing assemblies. The newest version, released last December, features an updated database that includes 500 new protein templates and enhanced accuracy. PROSPECT, the Protein Structure Prediction and Evaluation Computer Toolkit from Bioinformatics Solutions (BSI) of Waterloo, Ontario, employs threading to predict three-dimensional protein structure by recognizing native-like folds within a sequence. 
According to ApoCom's president, Robin Zimmer, the company's newest package, called Excavator (EXpression data Clustering Analysis VisualizATiOn Resource), is a gene expression database analysis application that allows the user to study functional relationships, clustering genes whose expression patterns correlate relative to time and/or altered conditions. Another offering, GeneHound, is a gene sequencing and identification program targeted for prokaryotic genomes, but which may be easily adapted to any genome. GrailEXP, a new, advanced version of the Grail Toolkit from ApoCom Genomics of Knoxville, Tenn., predicts both protein-coding genes and functional elements. This Windows-compatible software connects to the Wisconsin Package to perform database searches with minimal data configuration, and replaces the soon-to-be discontinued OMIGA, whose users will be offered a discounted upgrade to DS Gene. 
This June, the company launched its newest sequence analysis software package, Discovery Studio (DS) Gene. Last September, Accelrys released a Mac OS X-compatible version of MacVector. The Macintosh-compatible MacVector also offers multiple sequence analysis options and includes the AssemblyLIGN module to assemble consensus sequences. The UNIX-based package supports Web-based access via SeqWeb, and has a relational database version that integrates with SeqStore, the company's Oracle-based data management and mining system. What follows is a round up of each company's latest offerings.ĬORPORATE COMPENDIUM San Diego-based Accelrys' widely known GCG Wisconsin Package houses over 100 sequence analysis applications. DoubleTwist, supplier of TwistTools, has closed, and despite repeated efforts, we were unable to reach Gentech, Biotechnix3d's supplier, to confirm either the status of that program, or even if the company is still in business (its Web site has been under construction for some time). New companies on our list include ApoCom Genomics, Bioinformatics Solutions, and Paracel. Since our last review, (1) the software market has changed somewhat. Recently, companies have worked to make these programs user-friendly enough for anyone to use, often including detailed user manuals, get-started tutorials, and step-by-step protocol guides. 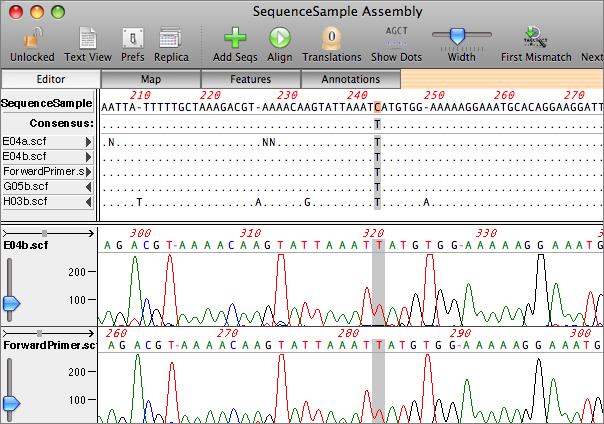
Most are also Web-enabled, allowing users to download data from remote databases. Researchers install most programs locally, on a desktop computer, but some packages are Web-accessible. Some sequence analysis programs include a broad range of these functions, whereas others include individual, task-specific modules smaller labs with specialized needs may find that purchasing particular modules better fits their budgets. Yet they can each perform some or all of the following functions: DNA and protein sequence entry, editing, annotation, analysis, and alignment primer identification map making contig assembly "in silico" cloning and gel prediction. These programs vary in terms of ease-of-use, power, and functionality. In the world of bioinformatics, the need to examine and manipulate large volumes of sequence data begat specialized computer software to handle these tasks. It is often said that necessity is the mother of invention.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
